
Karl Glastad
Assistant Professor
PhD
Research Active
Now accepting:
PhD students
Please email with inquiries.
Postdoctoral Applicants
Please email with inquiries.
- Office Location
- 306 Hutchison
- Telephone
- (585) 275-4891
- Web Address
- Website
Office Hours: By appointment
Research Overview
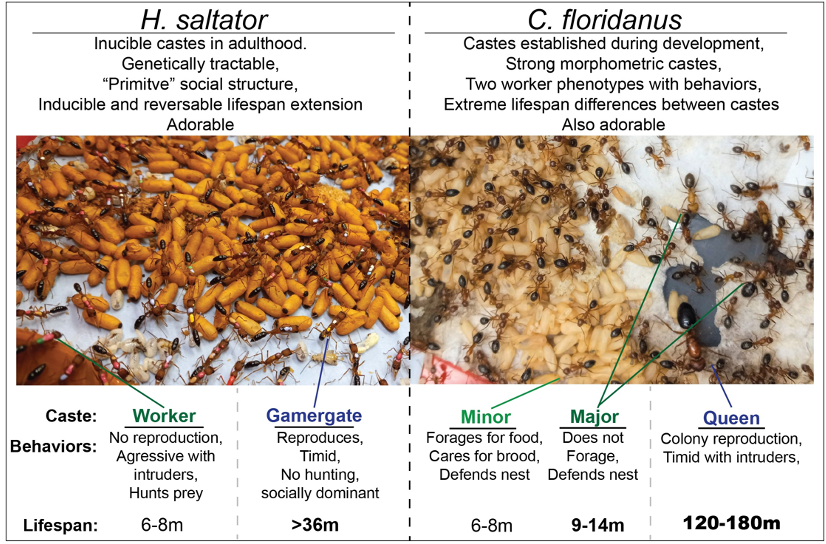

Ants live in sophisticated societies in which morphologically and behaviorally distinct types of individuals (castes) arise among members of closely-related families (colonies), and because of this, production of such castes is not genetic, but environmentally and epigenetically induced. Importantly, different castes typically exhibit often-extreme differences in behavior, physiology, and lifespan, which can be up to 20-fold longer in queens compared to workers Our lab employs two complementary ant systems as models to understand conserved and novel features governing the regulation of plasticity of behavior and lifespan.
Harpegnathos saltator: represents an ant with a simple social structure, wherein, if the queen is absent, a subset of workers can assume the reproductive roles within the colony as “gamergates” (GG). This worker-gamergate transition (W-GG) leads to dramatic physiological and behavioral changes, including a rapid change in behavior, reproductive physiology, and a >5-fold extension of lifespan. Further, GGs can be easily reverted back to the worker state, resulting in a reversion of behavior and physiology to the worker state, as well as a rapid loss of the extended lifespan seen in GG. This, along with the ability to perform transgenic manipulation, makes H. saltator an exception primary system within which to study plasticity of lifespan, as well as behavior and physiology.
Camponotus floridanus: provides an exceptional counterpoint to H. saltator as a more “advanced” ant, living in colonies with thousands of siblings and two worker castes (Minor and Major). Importantly, along with exhibiting different lifespans, Major and Minor workers perform distinct tasks, with Minor workers caring for brood and foraging, and Major workers defending the colony. Furthermore, while neither worker caste’s lifespan exceeds 1.5 years, C. floridanus queens possess a lifespan of ~20 years, exceeding the median lifespan of many mammals.
Our work involves a combination of in vivo work in ants, ectopic expression in D. melanogaster for quick screening, transient and transgenic perturbation in ants, and in vitro testing in ant neuronal cultures, as well as cell lines in fly and mouse.
The molecular, cellular, and organism-level causes and consequences of lifespan and age-related decline
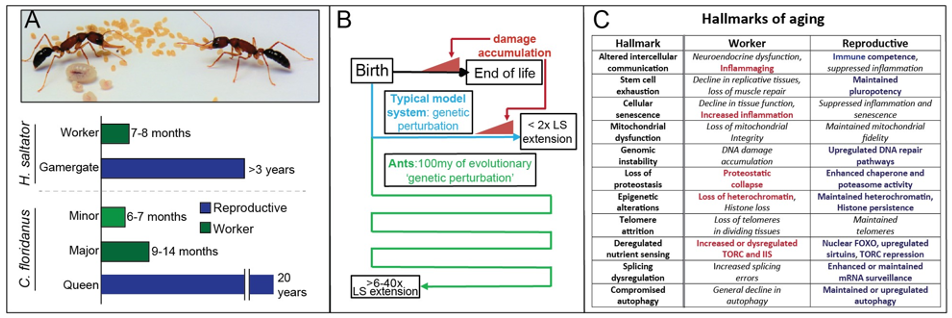
Aging is a complex syndrome associated with end-of-life (EoL) decline in organismal function associated with an accumulation of cellular and organismal damage, dysregulated signaling, and declines in gene regulatory fidelity. While much work has been done in conventional model systems to understand aging, the complex, multifaceted nature of age-related decline at EoL seen across the tree of life has complicated inferences and warrants alternative model systems to understand this complex process.
One of the main foci of our lab is to utilize ants as a model to understand stress response, lifespan, and the negative health consequences of aging. Ants provide an exceptional and largely unexplored system for this: unlike in conventional solitary organisms, in eusocial systems long lifespan of reproductives is evolutionarily advantageous. This has resulted in several features that make ants particularly promising for studies of aging. For one, because of these changes in evolutionary pressures across 100my of ant evolution, natural selection has performed all the ‘genetic perturbation’ required to accomplish drastic extension of reproductive ant lifespan (Panel A), something hampered in conventional systems by the highly-complex nature of age-related decline (Panel B). Furthermore, not only do ants possess lifespan differences that eclipse those seen between current species-level comparisons, they accomplish these differences between highly-related members of the same family via gene regulatory and epigenetic mechanisms.
This means that not only do ants provide an amazing system within which to understand extreme differences in lifespan, they provide a system where all of this is accomplished via alterations in gene regulation, not changes to the genome. This means that findings from studying ant lifespan plasticity are arguably more amenable to translational insight, as they provide mechanisms to alter lifespan and mitigate negative health consequences of aging via gene regulation and epigenetic perturbation.
Our prior work has identified a novel HSF2-like transcription factor up-regulated in long-lived reproductive workers in H. saltator, which is associated with enhanced heat stress-induced death and heat shock protein (HSP) up-regulation. However proteostasis (maintenance of the healthy proteome) is only one factor that declines with age. Indeed, age-related decline in cellular, tissue, and organism-level fitness is associated with a myriad of dysfunctions , often termed the “hallmarks of aging”, due to their consistency across Metazoa.
Preliminary data collected by our lab suggest that many, if not all of these “hallmarks” are being mitigated in long-lived reproductive ants (Panel C). Our lab seeks to build upon these data to understand:
- How ants circumvent each of these negative outcomes of aging at the molecular, cellular, and organismal levels,
- How reproductive ants are able to attenuate metabolism to accomplish the metabolically-demanding task of enormous egg production without the negative lifespan effects of such a metabolic state seen in solitary organisms,
- How this collective program of lifespan extension is orchestrated in the context of the neurohormonal signals that establish these castes,
- The convergent and novel mechanisms employed to regulate lifespan across disparate eusocial systems such as Termites (where queens live >30 years) as well as the Naked Mole Rat (Heterocephalus glaber), a mammalian eusocial system.
The epigenome, hormones, and brain microenvironment: Behavioral plasticity
 The ability to respond dynamically to changes in the environment is a hallmark of complex life. One of the major components of this in advanced multicellular organisms is modulation of behavior in response to environmental stimuli. Ants represent an outstanding system from which to study mechanisms governing behavioral adaptation, due to the production of multiple alternative behavioral states (castes) which are amenable to experimental manipulation. Previous work by us and others have established a strong experimental basis for inducing and understanding the molecular and neuronal bases for these behavioral states and shifts between them in both H. saltator and C. floridanus.
The ability to respond dynamically to changes in the environment is a hallmark of complex life. One of the major components of this in advanced multicellular organisms is modulation of behavior in response to environmental stimuli. Ants represent an outstanding system from which to study mechanisms governing behavioral adaptation, due to the production of multiple alternative behavioral states (castes) which are amenable to experimental manipulation. Previous work by us and others have established a strong experimental basis for inducing and understanding the molecular and neuronal bases for these behavioral states and shifts between them in both H. saltator and C. floridanus.
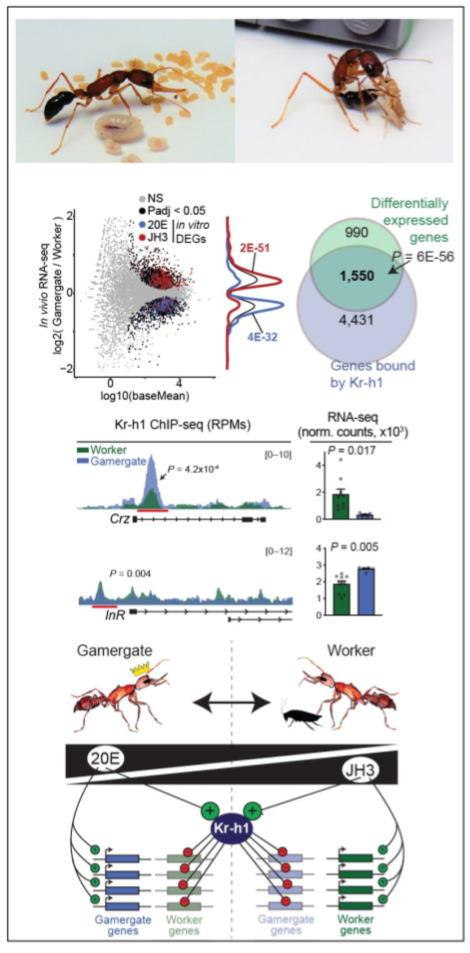
Neurohormonal basis for reproductive caste in H. saltator: Previously, in collaboration with Janko Gospocic (UTSW) we demonstrated that two hormones determine the worker-gamergate axis: Juvenile Hormone (JH3) signaling is associated with the worker state while Ecdysone (20E) is associated with an active reproductive status. We further found that the repressor Kr-h1 is involved in maintaining both of these, via repression of genes normally active only in the alternative caste.
We now seek to understand the mechanistic basis of this context-specific repression via Kr-h1, with a focus on nuclear hormone receptors, their binding of Kr-h1, and how these localize to alternative sets of genes depending on the dominant hormone for a given caste. This project will also seek to contextualize both the downstream targets, as well as nuclear hormone receptors in a mammalian context, to identify conserved and novel pathways associated with reproductive activation, social dominance, and appetitive responses.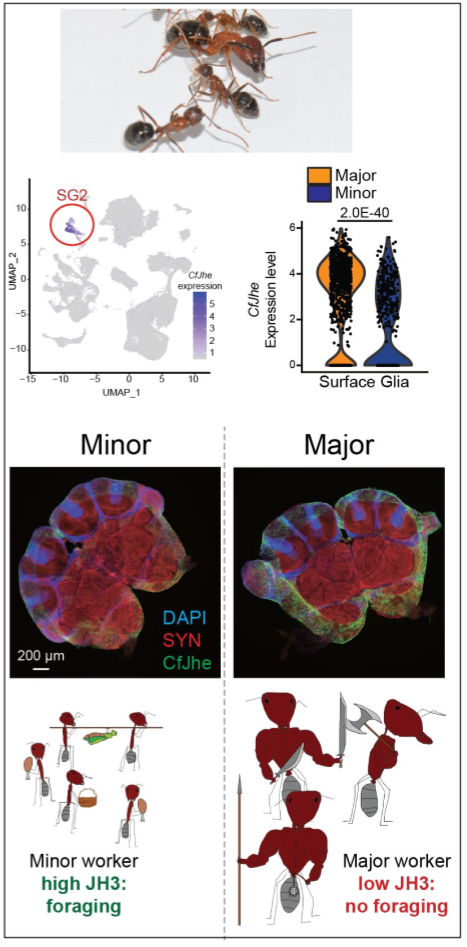
Origin, convergence, and regulation of the blood-brain barrier as a regulatory hub for behavioral plasticity: Recently, we demonstrated that one of the defining behavioral difference between Minor and Major workers in C. floridanus, foraging behavior, is controlled by differential degradation of the hormone Juvenile Hormone (JH3) by the workers’ blood-brain barrier (BBB). In Major workers which rarely forage, the BBB expresses high levels of a copy of Juvenile Hormone Esterase (Jhe, CfJhe hereafter), which unlike the copy in D. melanogaster, is retained in the cytoplasm and exclusively expressed in surface glia (cell of the insect BBB). In Minor workers, CfJhe is much more lowly expressed, leading to increased JH3 entry into the brain, and subsequently, foraging.
This unexpected and exciting finding opens the door to several projects related to understanding where this mechanism first originated in ants, if such mechanisms evolved repeatedly (convergent evolution) in ants with independent origins of multiple worker types, and if such hormone-gating by the BBB occurs in other taxa outside of Formicidae (ants).
Research Interests
- Gene regulation, hormonal signalling, molecular and cellular determinants of aging and lifespan; Mechanisms and evolution of plasticity; Arthropod genomics; Video games
Selected Publications
- Ju, L.*, Glastad, K.M.*^, Sheng, L., Gospocic, J., Kingwell, C.J., Davidson, S.M., Kocher, S.D., Bonasio, R., and Berger, S.L.^ (2023). Hormonal gatekeeping via the blood-brain barrier governs caste-specific behavior in ants. Cell, (in press). 10.1016/j.cell.2023.08.002. *Authors contributed equally; ^Co-corresponding authors link
- Glastad, K. M., Roessler, J., Gospocic, J., Bonasio, R., & Berger, S. L. (2023). Long ant life span is maintained by a unique heat shock factor. Genes & development, 37(9-10), 398-417. link
- Kee, J.^, Thudium, S.^, Renner, D. M.*, Glastad, K.M.*, Palozola, K., Zhang, Z., Li, Y., Lan, Y., Cesare, J., Poleshko, A., Kiseleva A.A., & Korb, E. (2022). SARS-CoV-2 disrupts host epigenetic regulation via histone mimicry. Nature, 610(7931), 381-388. ^*Authors contributed equally link
- Gospocic, J.*, Glastad, K. M.*, Sheng, L., Shields, E. J., Berger, S. L., & Bonasio, R. (2021). Kr-h1 maintains distinct caste-specific neurotranscriptomes in response to socially regulated hormones. Cell, 184(23), 5807-5823. *Authors contributed equally link
- Glastad, K. M., Ju, L., & Berger, S. L. (2021). Tramtrack acts during late pupal development to direct ant caste identity. PLoS Genetics, 17(9), e1009801. link
- Sheng, L.*, Shields, E. J.*, Gospocic, J., Glastad, K. M., Ratchasanmuang, P., Berger, S. L., Raj, A., Little, S., & Bonasio, R. (2020). Social reprogramming in ants induces longevity-associated glia remodeling. Science Advances, 6(34), eaba9869. *Authors contributed equally link
- Glastad, K. M.*, Graham, R. J.*, Ju, L., Roessler, J., Brady, C. M., & Berger, S. L. (2020). Epigenetic regulator CoREST controls social behavior in ants. Molecular Cell, 77(2), 338-351. *Authors contributed equally link
- Rehan, S. M.*, Glastad, K. M.*, Steffen, M. A., Fay, C. R., Hunt, B. G., & Toth, A. L. (2018). Conserved genes underlie phenotypic plasticity in an incipiently social bee. Genome biology and evolution, 10(10), 2749-2758. *Authors contributed equally link
- Gospocic, J., Shields, E. J., Glastad, K. M., Lin, Y., Penick, C. A., Yan, H., Mikheyev, A.S., Linksvayer, T.A., Garcia, B.A., Berger, S.L., Liebig, J., Reinberg, D., & Bonasio, R. (2017). The neuropeptide corazonin controls social behavior and caste identity in ants. Cell, 170(4), 748-759. link
- Glastad, K. M., Arsenault, S. V., Vertacnik, K. L., Geib, S. M., Kay, S., Danforth, B. N., Rehan, S.M., Linnen, C.R., Kocher, S.D., & Hunt, B. G. (2017). Variation in DNA methylation is not consistently reflected by sociality in Hymenoptera. Genome Biology and Evolution, 9(6), 1687-1698. link
- Glastad, K. M., Gokhale, K., Liebig, J., & Goodisman, M. A. (2016). The caste-and sex-specific DNA methylome of the termite Zootermopsis nevadensis. Scientific Reports, 6(1), 37110. link
- Glastad, K. M., Goodisman, M. A., Yi, S. V., & Hunt, B. G. (2016). Effects of DNA methylation and chromatin state on rates of molecular evolution in insects. G3: Genes, Genomes, Genetics, 6(2), 357-363. link
- Rehan, S. M.*, Glastad, K. M.*, Lawson, S. P., & Hunt, B. G. (2016). The genome and methylome of a subsocial small carpenter bee, Ceratina calcarata. Genome biology and evolution, 8(5), 1401-1410. *Authors contributed equally link
- Glastad, K. M., Hunt, B. G., & Goodisman, M. A. (2015). DNA methylation and chromatin organization in insects: insights from the ant Camponotus floridanus. Genome biology and evolution, 7(4), 931-942. link
- Glastad, K. M.*, Hunt, B. G.*, Yi, S. V., & Goodisman, M. A. (2014). Epigenetic inheritance and genome regulation: is DNA methylation linked to ploidy in haplodiploid insects?. Proceedings of the Royal Society B: Biological Sciences, 281(1785), 20140411. *Authors contributed equally link
