
Ning Wang
Assistant Professor
PhD
Research Active
Now accepting:
PhD students
Please email with inquiries.
- Office Location
- 436 Hutchison
- Telephone
- (585) 275-3890
- Web Address
- Website
Office Hours: By appointment
Research Overview
The major criterion distinguishing eukaryotes from prokaryotes is the presence of organelles in the former. An organelle is a subcellular structure that has one or more specific jobs to perform in the cell, much like an organ does in the body. Eukaryotic cells can rapidly adjust the abundance, size and shape of their organelles according to physiological needs. This remarkable capacity for adaptation enables cells to maintain homeostasis during stress, differentiation and disease. We are interested in the mechanisms cells use to ensure organelle homeostasis. We study the following two processes: 1) organelle biogenesis, including morphology generation, organelle inheritance during cell division, protein import and lipid transfer during organelle de novo synthesis 2) organelle degradation, particularly through the receptor-mediated selective autophagy pathway.
Organelle biogenesis
Organelle biogenesis is the process by which new organelles are made. Organelle biogenesis includes the process through which cellular organelles are split between daughter cells during mitosis; this process is called organelle inheritance. Membrane-bound organelles can also be generated through de novo synthesis where proteins and lipids both need to be inserted under precise control. Organelles often have their unique morphology that contributes to the specific function, often in ways that are poorly understood. We are particularly interested in the biogenesis of endoplasmic reticulum (ER), as it is the largest organelle in most eukaryotic cells and serves many essential cellular roles such as protein synthesis, lipid metabolism and calcium storage, etc. Currently we are asking questions such as: How do cells shape ER? How do they adjust ER size and function to physiological demand? How are ER inherited during cell division?
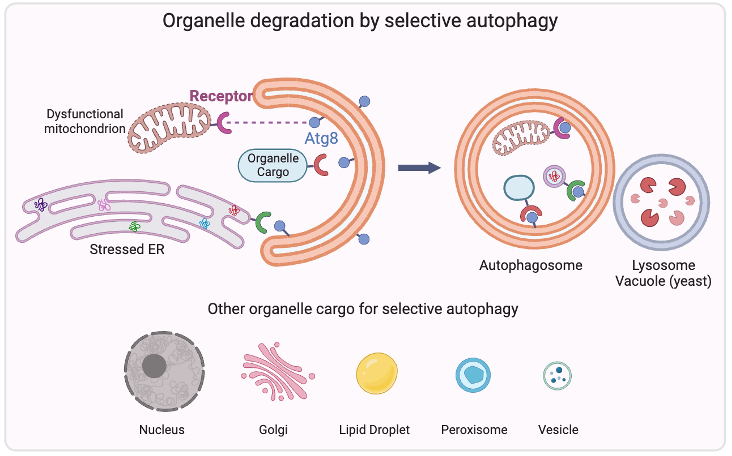
Organelle degradation
Autophagy is a highly conserved physiological process that degrades excess or damaged organelles, large protein aggregates and invading pathogens via the lysosomal system (the vacuole in plants and yeast). Autophagy can be non-selective bulk degradation, or selective. Selective autophagy recycles specific organelles, such as mitochondria and ER through receptor-mediated cargo selection according to the physiological needs. This capability makes selective autophagy a major process in maintaining organelle homeostasis, and the dysfunction of selective autophagy underlines many human diseases, including neurodegeneration, cancer, and metabolic disorders. We’re interested in studying the “selective” mechanism, and the current research focus is to identify specific autophagy receptors for different organelles such as nucleus, golgi, lipid droplets and peroxisomes.
Research Interests
- Cell biology, biochemistry, and molecular genetics
- Organelle biogenesis
- Organelle degradation through selective autophagy
- Organelle aging and neurodegeneration
Selected Publications
- Wang, N., Y. Shibata, J.A. Paulo, S.P. Gygi, and T.A. Rapoport. (2023). A conserved membrane curvature-generating protein is crucial for autophagosome formation in fission yeast. Nat Commun 14, 4765.
- Wang, N., L.D. Clark, Y. Gao, M.M. Kozlov, T. Shemesh, and T.A. Rapoport. (2021). Mechanism of membrane-curvature generation by ER-tubule shaping proteins. Nat Commun 12, 568.
- Wang, N. and T.A. Rapoport. (2019). Reconstituting the reticular ER network - mechanistic implications and open questions. J Cell Sci 132(4):jcs227611.
- Wang, N., I-J. Lee, G. Rask, and J.-Q. Wu. (2016). Roles of the TRAPP-II Complex and the Exocyst in Membrane Deposition during Fission Yeast Cytokinesis. PLoS Biol 14(4): e1002437.
- Wang, N., M. Wang, Y.-H. Zhu, T.W. Grosel, D. Sun, D.S. Kudryashov, and J.-Q. Wu. (2015). The Rho-GEF Gef3 interacts with the septin complex and activates the GTPase Rho4 during fission yeast cytokinesis. Mol Biol Cell 26(2):238-55.
- Wang, N., L. Lo Presti, Y.-H. Zhu, M. Kang, Z. Wu, S.G. Martin, and J.-Q. Wu. (2014). The novel proteins Rng8 and Rng9 regulate the myosin-V Myo51 during fission yeast cytokinesis. J Cell Biol 205(3):357-75.
