
Daven C. Presgraves
Shirley Cox Kearns Professor of Biology
Professor of Biology
PhD
Research Active
Now accepting:
PhD students
Please email with inquiries.
- Office Location
- 481 Hutchison
- Telephone
- (585) 275-0925
- Web Address
- Website
Office Hours: By appointment
Research Overview
We study the evolution & consequences of selfish genetic elements. Recurrent invasion & innovation by selfish genetic elements and suppression & counter-innovation by hosts are major, if still unappreciated, drivers of genome evolution. Our lab combines classical, molecular, and evolutionary genetics & genomics to study how selfish genetic elements— such as transposons and meiotic drivers— affect speciation, sex chromosome evolution, and meiotic recombination.
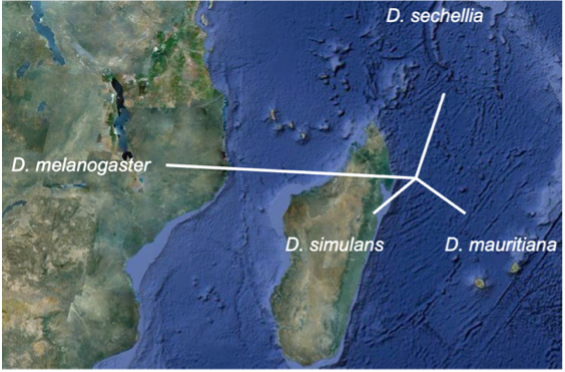
Hybrid incompatibility. Historically, one of the major gaps in speciation genetics research was the limited number of hybrid incompatibility genes identified at the molecular level. This situation began to change ~15 years, with a surprising discovery: hybrid incompatibility genes evolve via recurrent positive selection as they mediate molecular arms races between hosts and their selfish genetic elements. Two hybrid incompatibility genes that we identified between Drosophila melanogaster and its sibling species interact with the host transposon surveillance machinery. In new work involving the much younger species of the D. simulans clade species— D. mauritiana, D. simulans, and D. sechellia— we have identified a direct role for transposons in hybrid incompatibility.
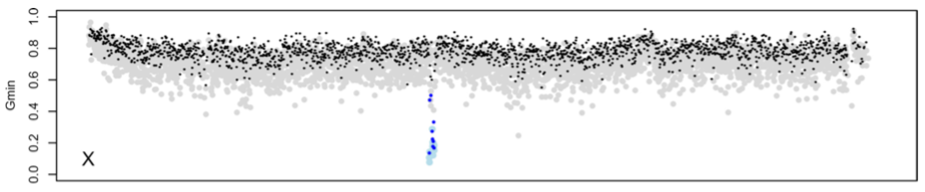
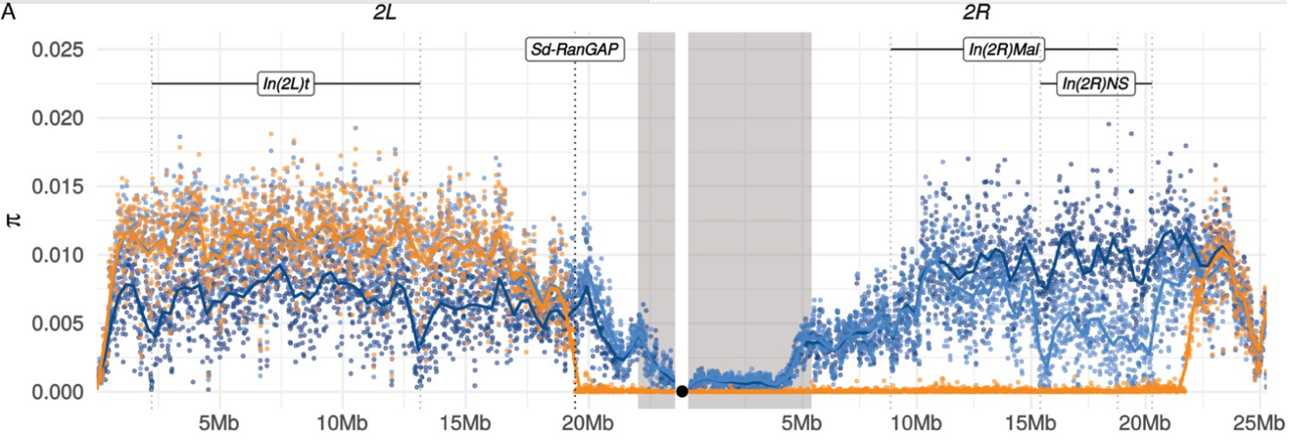
Sex chromosomes and speciation in flies. Sex chromosomes play a special role in speciation, as evidenced by three major “rules” of speciation: Haldane’s rule, the large X-effect, and the tendency for X and Y (Z and W) chromosomes to show reduced interspecific gene flow. Our genome-wide introgression experiments and population genomics analyses in the D. simulans clade species show that the rapid accumulation of hybrid male sterility factors on the X can explain Haldane’s rule, the large X-effect, and its reduced propensity for natural introgression. This work has revealed a surprising interplay between gene flow, hybrid male sterility, and meiotic drive.

Neo-sex chromosomes and speciation in birds. In collaboration with Al Uy’s lab, we are studying the evolutionary genomics of neo-sex chromosomes and speciation in oceanic island birds. About a century ago, two species of Myzomela honeyeaters established secondary contact and began to hybridize regularly. We are now investigating how gene flow, natural selection, and behavior have shaped the amount, direction, and distribution of genomic admixture.
Meiotic drive. We study the genetics, molecular biology, evolution, population dynamics, and genomic consequences of meiotic drive elements— selfish genes that enhance their own transmission by distorting fair Mendelian inheritance— in two systems. The autosomal Segregation Distorter (SD) of Drosophila melanogaster is a classic selfish supergene that comprises multiple, linked, epistatically interacting loci and suppressors of recombination. And the recently evolved, cryptic sex-ratio drive systems of D. simulans clade species each comprises multi-copy sets if X-linked drivers and specialized endogenous small RNA-encoding autosomal suppressors.
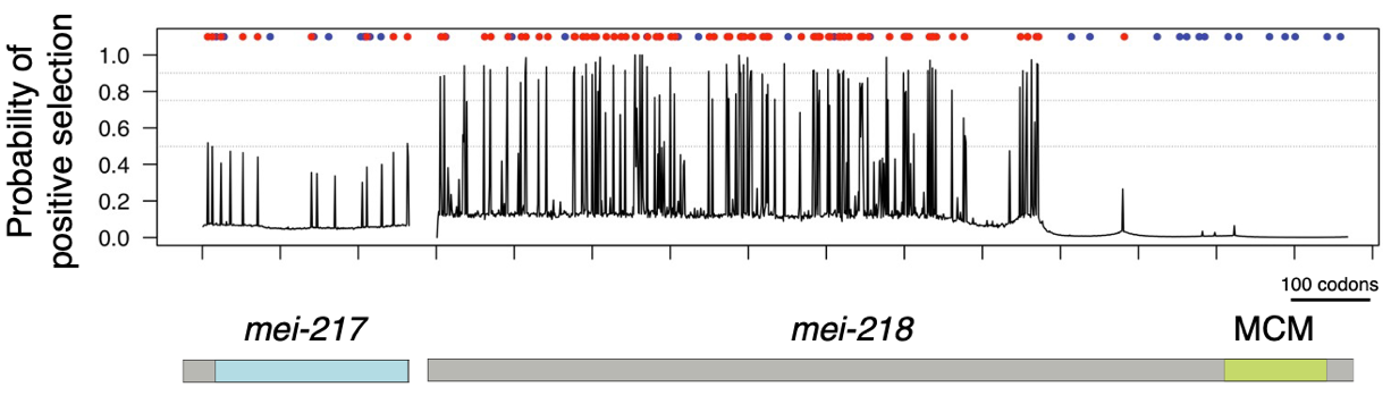
The evolution of meiotic recombination rate & patterning. Crossing over between homologous chromosomes during meiosis repairs DNA double-strand breaks, ensures proper segregation, and shapes the genomic distribution of genetic variability. We have also shown that recombination enhances the efficacy of natural selection in Drosophila genomes. But why species with identical karyotypes evolve big differences in rates of crossing over is unclear. We found that a single meiosis gene with a history of positive selection mediates most of the species difference in crossing over between D. melanogaster and D. mauritiana. Optimal crossover rates may evolve continuously to balance the benefits of recombination against the ever-shifting risk of ectopic exchange posed by transposons.
Research Interests
- Evolutionary genetics
- Speciation genetics
- Molecular population genetics
- Selfish gene complexes
Selected Publications
- Navarro-Dominguez, B., C.-H. Chang, C.L. Brand, C.A. Muirhead, D.C. Presgraves and A.M. Larracuente. 2022.Epistatic selection on a selfish SegregationDistorter supergene: drive, recombination, and genetic load. eLife11:e78981.
- Muirhead, C.A. and D.C. Presgraves. 2021.Satellite DNA-mediated diversification of a sex-ratio meiotic drive gene family in Drosophila. Nature Ecology & Evolution 5:1604-1612.
- Presgraves, D.C. and C.D. Meiklejohn. 2021.Hybrid sterility, genetic conflict and complex speciation: lessons from the Drosophila simulans clade specieFrontiers in Genetics 12: 669045.
- Meiklejohn, C.D., E.L. Landeen, K.E. Gordon, T. Rzatkiewicz, S.B. Kingan, A. Geneva, J.P. Vedanayagam, C.A. Muirhead, D. Garrigan, D.L. Stern, and D.C. Presgraves. 2018. Gene flow mediates the role of sex chromosome meiotic drive during complex speciation. eLife 7: e35468.
- Presgraves, D.C. 2018.Evaluating genomic signatures of “the large X-effect” during complex speciation. Molecular Ecology 27:3822-3830.
- Brand, C.L., M.V. Cattani, S.B. Kingan, E.L. Landeen and D.C. Presgraves. 2018. Molecular evolution at a meiosis gene mediates species differences in crossing over. Current Biology 28:1289-1295.
- Landeen, E.L., C.A. Muirhead, L. Wright, C.D. Meiklejohn and D.C. Presgraves. 2016.Sex chromosome-wide transcriptional suppression and compensatory cis-regulatory evolution mediate gene expression in the Drosophila male germline. PLoS Biology 14:1002499.
- Lindholm, A.K., K.A. Dyer, R.C. Firman, L. Fishman, W. Forstmeier, L. Holman, H. Johannesson, U. Knief, H. Kokko, A.M. Laraccuente, A. Manser, C. Montchamp-Moreau, V.G. Petrosyan, A. Pomiankowski, D.C. Presgraves, L.D. Safronova, A. Sutter, R.L. Unckless, R. Verspoor, N. Wedell, G.S. Wilkinson, T.A.R. Price. 2016.The ecology and evolutionary dynamics of meiotic drive. Trends in Ecology and Evolution 31: 315-326.
- Muirhead, C.A. and D.C. Presgraves. 2015. Hybrid incompatibilities, local adaptation, and the genomic distribution of natural introgression between species. American Naturalist 187: 249-261.
- Tang, S. and D.C. Presgraves 2015 Lineage-specific evolution of the complex Nup160hybrid incompatibility in Drosophila.Genetics 200: 1245-1254.
- Brand, C.L., A.M. Larracuente and D.C. Presgraves 2015 Origin, evolution, and population genetics of the selfish Segregation distorter gene duplicaion in European and African populations of Drosophila melanogaster. Evolution 69: 1271-1283.
- Garrigan, D., S.B. Kingan, A.J. Geneva, J.P. Vedanayagam and D.C. Presgraves 2014 Genome diversity and divergence in Drosophila mauritiana: multiple signatures of faster X evolution.Genome Biology & Evolution 6: 2444-2458.
- Meiklejohn, C.D. and D.C. Presgraves 2012 Little evidence for demasculinization of the Drosophila X chromosome among genes expressed in the male germline.Genome Biology and Evolution 4: 895-904.
- Larracuente, A.L. and D.C. Presgraves 2012 The selfish Segregation Distorter complex of Drosophila melanogaster.Genetics 192: 33-53.
- Garrigan, D., S. Kingan, A. Geneva, K. Thornton, P. Andolfatto, A.G. Clark, and D.C. Presgraves 2012 Genome sequencing reveals complex speciation of the Drosophila simulans clade.Genome Research 22: 1499-1511.
- Gérard, P.R. and D.C. Presgraves. 2012. Abundant genetic variability in Drosophila simulans for hybrid female lethality in interspecific crosses to D. melanogaster.Genetics Research 94: 1-7.
- Meiklejohn, C.D., E.L. Landeen, J.M. Cook, S.B. Kingan and D.C. Presgraves. 2011. Sex chromosome-specific regulation in the Drosophila male germline but little evidence for chromosomal dosage compensation or meiotic inactivation. PLoS Biology 9: e1001126.
- Presgraves, D.C. 2010. Darwin and the origin of interspecific genetic incompatibilities. American Naturalist 176: S45-S60.
- Presgraves, D.C. 2010. The molecular evolutionary basis of species formation. Nature Reviews Genetics 11: 175-180.
- Presgraves, D.C. and S.V. Yi. 2009. Doubts about complex speciation between humans and chimpanzees. Trends in Ecology & Evolution 24: 533-540.
- Presgraves, D.C., P.R. Gérard, A. Cherukuri and T.W. Lyttle. 2009. Large-scale selective sweep among Segregation Distorter chromosomes in African populations of Drosophila melanogaster. PLoS Genetics 5: e1000463.
- Cattani, M.V. and D.C. Presgraves. 2009. Genetics and lineage-specific evolution of a lethal hybrid incompatibility between Drosophila mauritiana and its sibling species. Genetics 181: 1545-1555.
- Tang, S. and D.C. Presgraves. 2009. Evolution of the Drosophila nuclear pore complex results in multiple hybrid incompatibilities. Science 323: 779-782.
- Presgraves, D. C. 2008.Sex chromosomes and speciation in Drosophila.Trends in Genetics.24: 336-343.
- Masly, J.P. and D.C. Presgraves. 2007. High-resolution genome-wide dissection of the two rules of speciation in Drosophila. PLoS Biology 5: 1890-1898.
- Presgraves, D.C. and W. Stephan. 2007. Pervasive adaptive evolution among interactors of the Drosophila hybrid inviability gene, Nup96. Mol. Biol. Evol. 24: 306-314.
- Presgraves, D.C. 2005. Recombination enhances protein adaptation in Drosophila melanogaster. Current Biology 15: 1651-1656.
- Presgraves, D.C., L. Balagopalan, S.A. Abmayr and H.A. Orr. 2003. Adaptive evolution drives divergence of a hybrid inviability gene between two species of Drosophila. Nature 243: 715-719.
- Presgraves, D.C. 2003. A fine-scale genetic analysis of hybrid incompatibilities in Drosophila. Genetics 163: 955-972.
- Presgraves, D.C. 2002. Patterns of postzygotic isolation in Lepidoptera. Evolution 56: 1168-1183.
- Betancourt, A.B. and D.C. Presgraves. 2002. Linkage limits the power of natural selection in Drosophila. PNAS 99: 13616-13620.
- Presgraves, D.C. and H.A. Orr. 1998. Haldane's rule is obeyed in taxa lacking a hemizygous sex. Science 282: 952-954.
