Nucleic Acids Res 1998 Dec 1;26(23):5394-401
Department of Biology, University of Rochester, Rochester, NY 14627, USA.
Sequence-specific transcription factors need to gain access to regulatory sequences in chromatin. Previous studies utilizing model systems have suggested many mechanisms involved in this process. It is unclear however how these findings relate to natural promoters. The Drosophila alcohol dehydrogenase ( Adh ) gene distal promoter is organized into an ordered nucleosome array before multiple transcription factors recognize their sites within this nucleosomal context and activate transcription. Here we used a purified in vitro system to study the binding of the ubiquitous Drosophila transcription factors Adf-1 and GAGA factor to the Adh distal promoter in chromatin. Several nucleosome core particles were assembled on 150 bp DNA fragments containing the Adh distal cis -acting elements in the natural promoter context but different DNA-histone environments. We found that the Adh distal promoter regulatory sequences can position nucleosomes in the same rotational setting as observed in vivo. In one particular nucleosome position, the wrapping of the Adf-1 and adjacent GAGA factor binding sitesaround the histone octamer creates a unique local DNA conformation. High-affinity but non-cooperative nucleosome binding of Adf-1 and GAGA factortherefore occurs, in contrast to the inhibition of Adf-1 and GAGA factor binding in other nucleosome positions. Thus, local histone-DNA sequence contact giving rise to a specific asymmetric nucleosome structure may play important roles in modulating the affinities of transcription factors for their nucleosomal sites.
PMID: 9826764, UI: 99045605
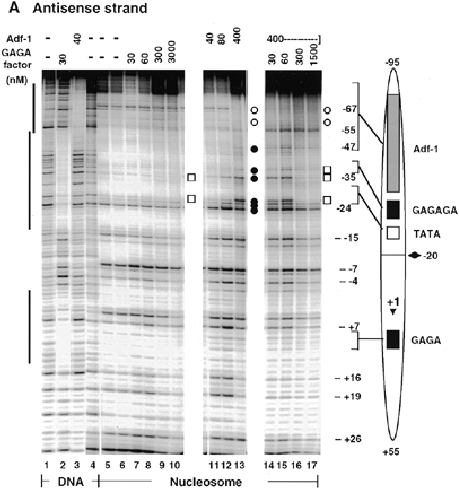
Sequence-specific binding of Adf-1 and GAGA factor to nucleosome I.
(A) DNase I footprinting of Adf-1 and GAGA factor on the antisense strand of DNA fragment I (-95 to +55) as free DNA (lanes 1-4) or nucleosome core particles (lanes 5-17).
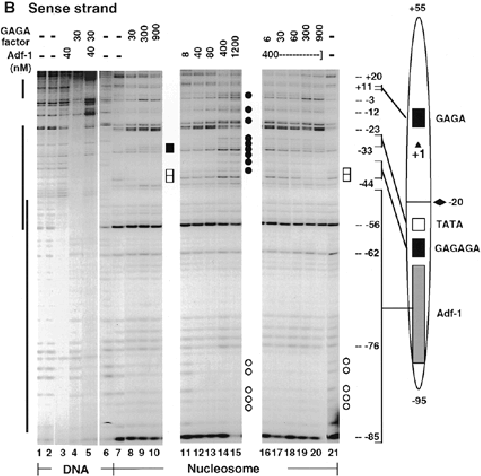
(B) DNase I footprinting of Adf-1 and GAGA factor on the sense strand of DNA fragment I (-95 to +55) as free DNA (lanes 1-6) or nucleosome core particles (lanes 7-21). Note that the binding conditions were equalized for DNA and nucleosome core particles (see article for details) which led to higher protein concentrations for footprinting detection than those in the DNase I footprint titration analysis. GAGA factor refers to either GAGA-519aa or GAGA-581aa since they have the same DNA and nucleosome binding properties individually or together in our experiments. Vertical bars on the left indicate the positions f footprints on naked DNA. The diagram of nucleosome I is shown to the right. Open squares, protection on the nucleosome structure by GAGA factor; open circles, protection by Adf-1; solid squares, hypersensitivity induced by GAGA factor; solid circles, hypersensitivity induced by Adf-1.


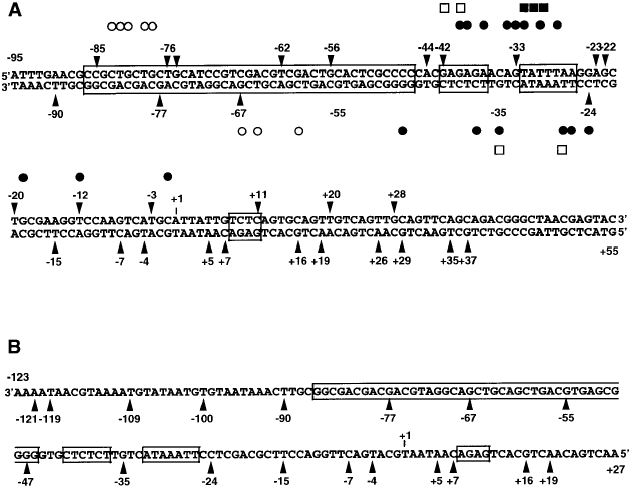
Summary of the DNase I footprinting data of nucleosome I (A) and nucleosome II (B).
The factor binding sites on naked DNA are boxed. Arrow heads indicate the DNase I cleavage sites on nucleosome core particles. Open and solid squares are as described above.